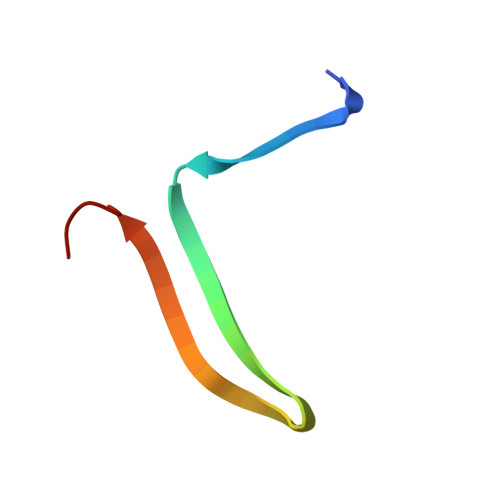
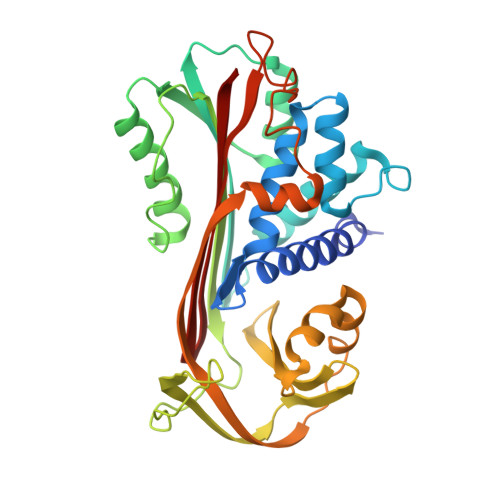
The S variant of human alpha 1-antitrypsin, structure and implications for function and metabolism.
Engh, R., Lobermann, H., Schneider, M., Wiegand, G., Huber, R., Laurell, C.B.(1989) Protein Eng 2: 407-415
- PubMed: 2785270 
- DOI: https://doi.org/10.1093/protein/2.6.407
- Primary Citation of Related Structures:  
7API, 8API, 9API - PubMed Abstract: 
The S variant of the human alpha 1-antitrypsin with E-264----V, is responsible for a mild alpha 1-antitrypsin deficiency quite common in the European population. S protein specifically cleaved at the susceptible peptide bond was crystallized and its crystal structure determined and refined to 3.1 A resolution. The S variant crystallizes isomorphous to the normal M variant. The difference Fourier electron density map shows the E----V change as outstanding residual density. In addition, small structural changes of the main polypeptide chain radiate from the site of mutation and affect parts far removed from it. By the mutation, internal hydrogen bonds and salt linkages of E-264 to Y-38 and K-487, respectively, are lost. They cause the far-reaching slight distortions and are probably related to the reduced thermal stability of the S mutant. They may also be responsible for slower folding of the polypeptide chain and the clinical symptoms of alpha 1-antitrypsin deficiency. In a theoretical study by molecular dynamics methods simulations of the M and S proteins were made and the results analysed with respect to structural and dynamic properties and compared with the experimental results. There is a significant correlation between experimental and theoretical results in some respects.
- Max-Planck-Institut für Biochemie, Martinsried, FRG.
Organizational Affiliation: